Researchers reveal the metabolic capacity of a termite gut protist and its associated bacteria using single-cell (meta)genomics
Scientists from the Faculty of Science CUNI-BIOCEV (Sebastian Treitli a Vladimír Hampl) in collaboration with researchers from University of British Columbia, sequenced the genomes of an isolated individual cell of the protist, Streblomastix strix, and at least eight different species or strains of bacterial symbionts which cover its surface. They used this genomic data to reconstruct the metabolism within this complex symbiotic system.
The coding capacity of individual components suggest that enzymes secreted by the bacteria drive the cellulose digestion resulting in simple sugars. Bacteria also have the capacity to fix nitrogen and synthesize most amino acids and cofactors. The protist likely uses simple sugars from cellulose degradation in its vicinity and may also acquire amino acids, vitamins and cofactors by occasional phagocytizing and digesting surface symbionts. Interestingly, most bacteria lack enolase, an enzyme essential for their core energy metabolism (glycolysis). Authors speculate that the protist complements the enolase activity and thus maintains the control over the bacterial component of the system.
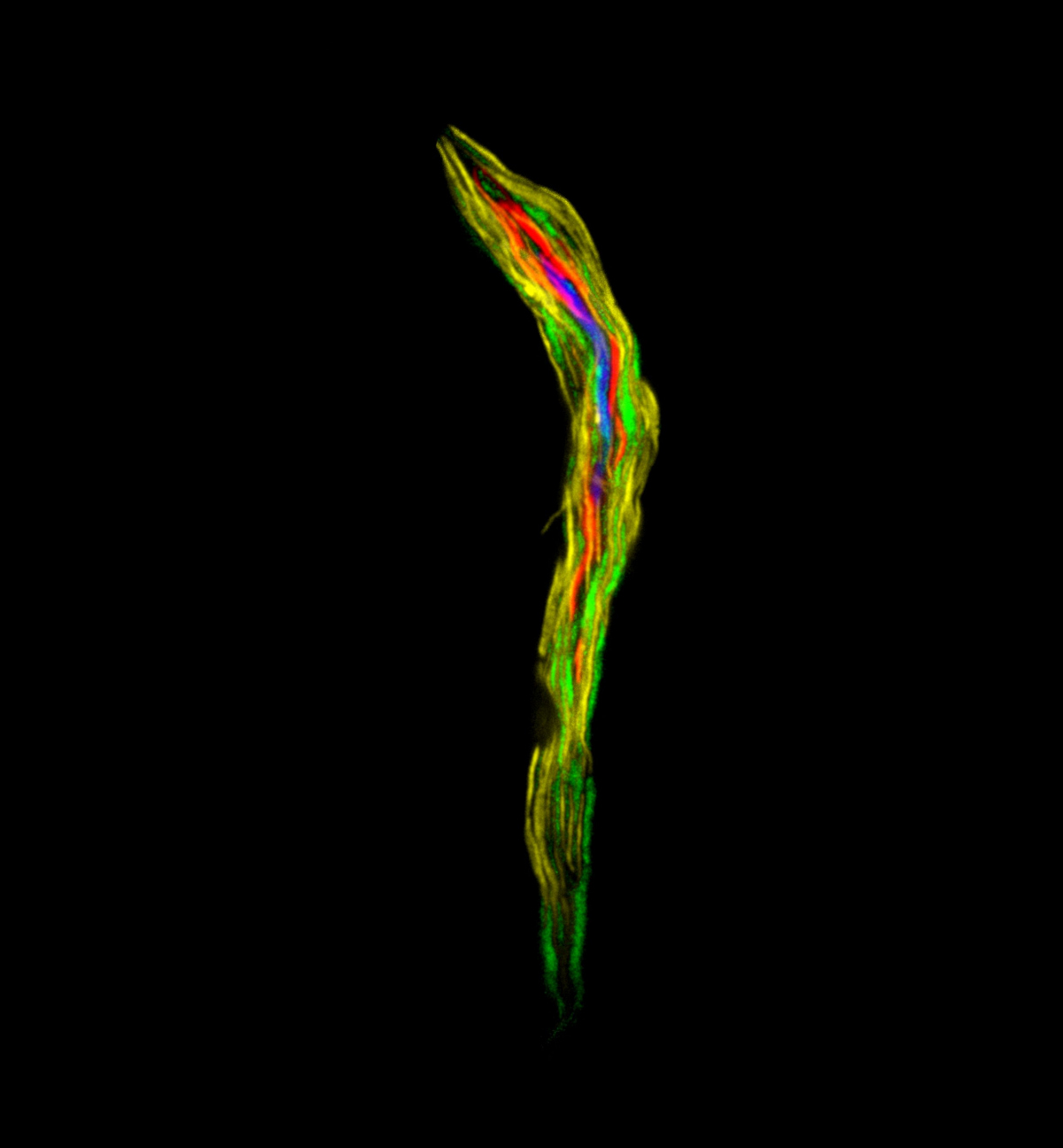
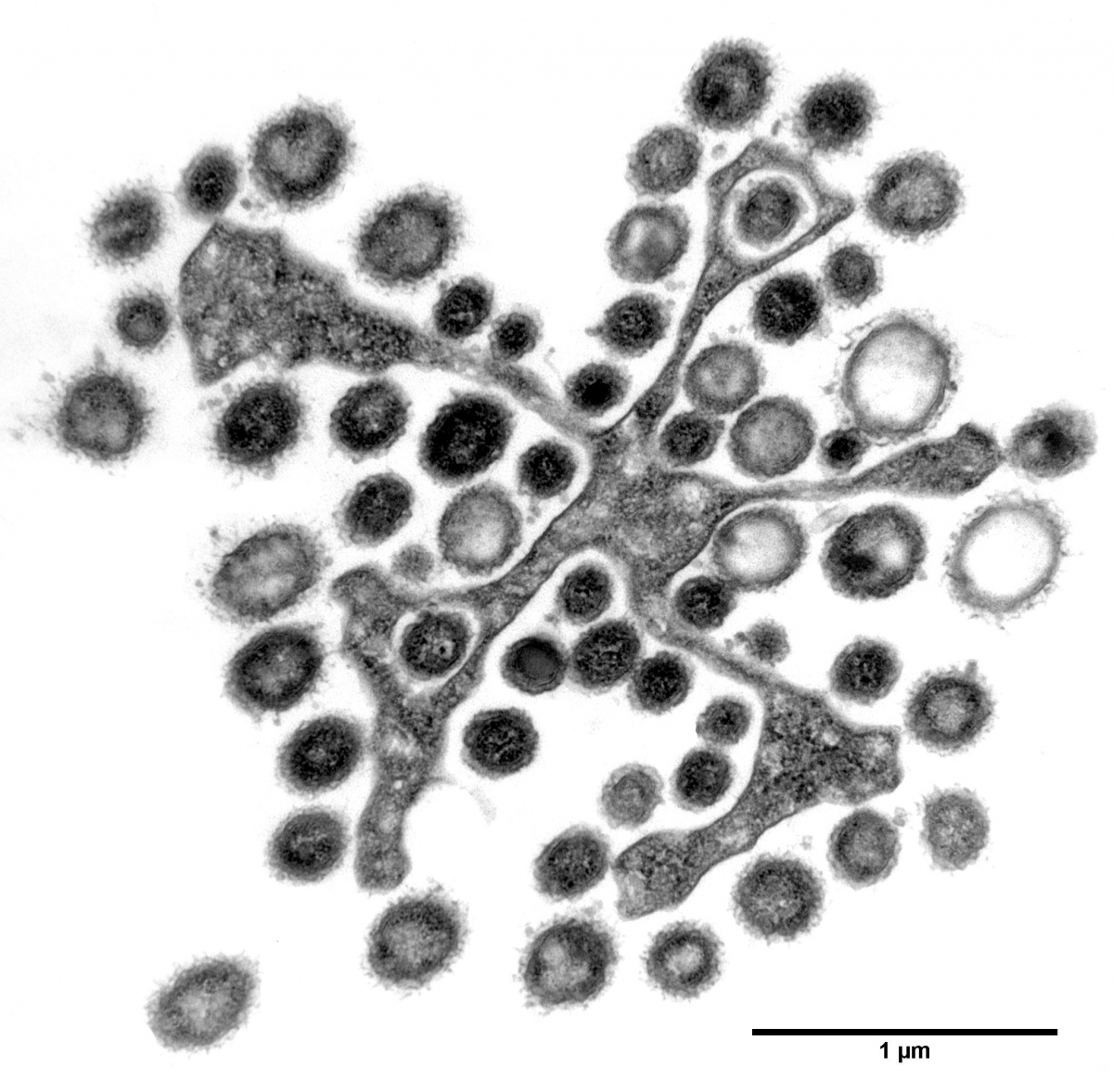
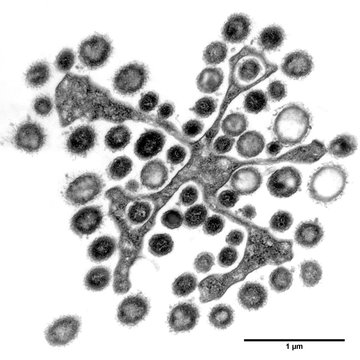
Lower termites form complex symbiotic relationships with bacteria and protists, which help them to digest wood. For the termites, this is an obligate relation, as loss of the microbes inhabiting their gut leads to their death. In the termite gut, many protists enter symbiotic relations with various bacteria which are located inside or on the surface of their cells, sometimes forming a spatially structured superorganism. This complex termite-protist-bacteria symbiosis is very poorly understood, the role of each member is unclear and will probably vary from case to case.
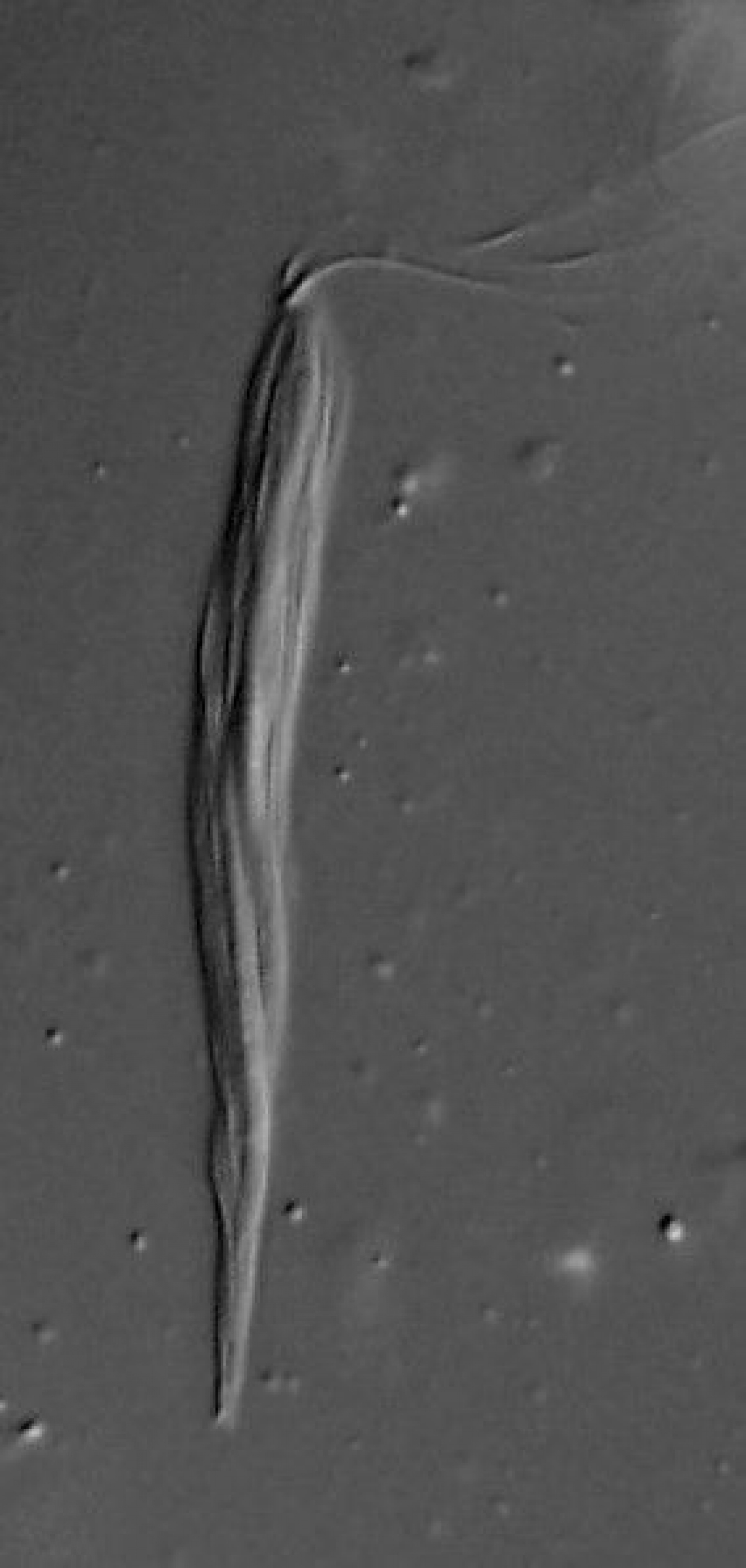
Sebastian C. Treitli, Martin Kolisko, Filip Husník, Patrick J. Keeling, Vladimír Hampl: Revealing the metabolic capacity of Streblomastix strix and its bacterial symbionts using single-cell metagenomics Proceedings of the National Academy of Sciences Sep 2019, 201910793; DOI: 10.1073/pnas.1910793116