About us
RESEARCH AREAS:
Bioactive compounds from bacteria, fungi and invertebrates
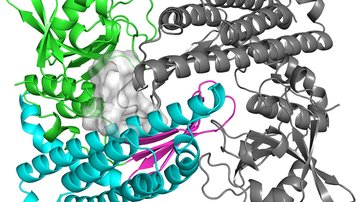
The structure and function of biotechnologically relevant enzymes
Antibiotic mode of action and mechanisms of resistance
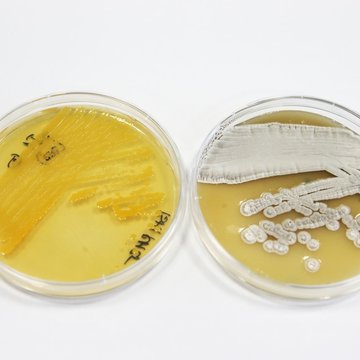
Characterization of bacterial human and envirnonmental communities
MAIN OBJECTIVES:
Determination of the structure and mechanism of action of natural biologically active compounds, understanding of their role in the environment as well as their interactions with target molecules
Characterization of the biosynthetic, biotransformational and biodegradational potential of bacteria, fungi and invertebrates, including the properties and structure of their enzymes
Study of the metabolic activity of microbial producers of natural substances in the environment using metagenomics, metatranscriptomics and metaproteomics.
CONTENT OF THE RESEARCH:
Gabriela Balíková Novotná
The problem of antibiotic resistance in gram-positive cocci is addressed using a complex approach. The function of resistance proteins and mechanisms of resistance development is studied in pathogens and antibiotic producers. The knowledge is then used for the development of drugs to overcome the existing resistance mechanisms and/or to be active against biofilms. The activity of new drugs, the mode of their action and possibility of antibiotic resistance development is assessed. Biosynthetic pathways of antibiotic production are also investigated and participating enzymes are characterized in terms of their function and structure.
Rubén López Mondéjar
The composition and ecology of communities of natural compound-producing microorganisms in the environment are studied with the use of metagenomics metatranscriptomics and metaproteomics. Functional screening and characterization leads to the recovery of microbial strains with biotechnological potential, their characterization using genome sequencing and proteomics and the exploration of the regulation of the production of natural, biologically-active compounds, proteins and genes. Novel free-living bacteria and fungi as well as those living in symbiotic relationships with other organisms (plants, insects, mammals) are targeted.
POTENTIAL FOR COOPERATION:
Testing the clinical potential of new antibacterial compounds. Exchange of research fellowships is welcome. Student positions at all levels are advertised. Cooperation with end-users of the biotechnologically-relevant environmental strains of microorganisms, their enzymes and genes.